_15.png)
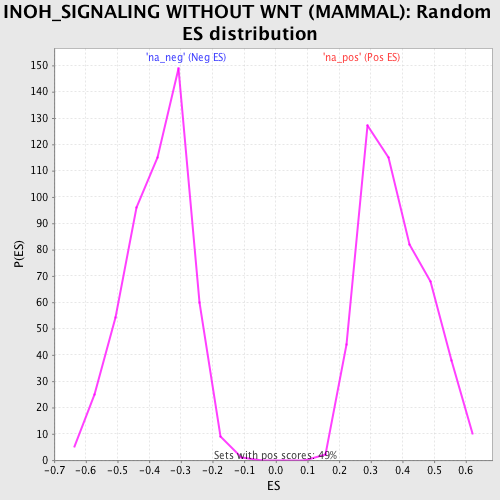
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | INOH_SIGNALING WITHOUT WNT (MAMMAL) |
| Enrichment Score (ES) | 0.70253795 |
| Normalized Enrichment Score (NES) | 1.8698429 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.043828476 |
| FWER p-Value | 0.269 |
_15.png)
| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TCF7L2 | 72 | 20.268 | 0.1829 | Yes | ||
| 2 | LEF1 | 176 | 14.378 | 0.3099 | Yes | ||
| 3 | TLE4 | 336 | 11.842 | 0.4104 | Yes | ||
| 4 | LRP5 | 863 | 7.384 | 0.4502 | Yes | ||
| 5 | TLE6 | 888 | 7.236 | 0.5156 | Yes | ||
| 6 | TLE3 | 1308 | 5.500 | 0.5437 | Yes | ||
| 7 | SMAD4 | 1551 | 4.811 | 0.5750 | Yes | ||
| 8 | CREBBP | 1702 | 4.447 | 0.6080 | Yes | ||
| 9 | AXIN2 | 1852 | 4.127 | 0.6380 | Yes | ||
| 10 | TCF7 | 2084 | 3.666 | 0.6593 | Yes | ||
| 11 | GSK3B | 2190 | 3.495 | 0.6859 | Yes | ||
| 12 | APC | 2416 | 3.120 | 0.7025 | Yes | ||
| 13 | DVL2 | 3177 | 2.221 | 0.6821 | No | ||
| 14 | BCL9 | 3470 | 1.955 | 0.6844 | No | ||
| 15 | DVL1 | 5512 | 0.813 | 0.5822 | No | ||
| 16 | NLK | 7210 | 0.198 | 0.4927 | No | ||
| 17 | FRAT2 | 7632 | 0.071 | 0.4707 | No | ||
| 18 | TCF3 | 8029 | -0.054 | 0.4499 | No | ||
| 19 | TLE2 | 11153 | -0.945 | 0.2907 | No | ||
| 20 | PYGO1 | 11913 | -1.170 | 0.2606 | No | ||
| 21 | DVL3 | 13281 | -1.604 | 0.2019 | No | ||
| 22 | CUL1 | 14455 | -2.118 | 0.1583 | No | ||
| 23 | AXIN1 | 14596 | -2.191 | 0.1710 | No | ||
| 24 | TLE1 | 16984 | -4.902 | 0.0878 | No |